I recently wrote a function that performs multiple linear hypothesis tests on a linear model object in R. As this is a common use of regression models, I am sure there are other easy-to-use packages out there, but my function works well (so feel free to use it).
The function is called test.lin(), and here is the code that you'll need to run to get it to work:
test.lin = function(R, co, model,...){
z = summary(model)
cov = z$cov.unscaled
beta = z$coefficients[,1]
g = nrow(R)
sig.sq = (z$sigma)^2
Rb.h = R%*%beta
diff = Rb.h-co
var.con = R%*%cov%*%t(R)
var.con.inv = solve(var.con)
W = t(diff)%*%var.con.inv%*%diff/(g*sig.sq)
df= model$df.residual
p.val= pf(W,g,df,lower.tail=F)
d = list(Rb.h, co, W, p.val)
names(d) = c("Prediction","Ho Value","Test Stat",
"P-value")
d
}
Once you run the above code, the function
test.lin() can be used to test multiple hypothesis of the form:
Ho: RBeta = coHa: RBeta !=coWhere R is a matrix and Beta is a vector of regression parameters. Each row of R defines a linear combination of regression parameters. Then, the function allows the user to specify the hypothesized value of each linear combination with the vector co.
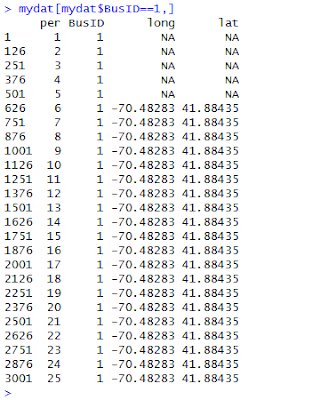
Here's an example of how to use the function. To use my code, you will want to
read in the data (found
here) and estimate a linear model called school.lm:
> ## ----------------------------- ##
> ## Test that beta1=beta2=beta3=0 ##
> ## ----------------------------- ##
>
> R1 = c(0, 1, 0, 0, 0)
> R2 = c(0, 0, 1, 0, 0)
> R3 = c(0, 0, 0, 1, 0)
>
> R = rbind(R1, R2, R3)
> co = matrix(c(0,0,0),nrow=3)
>
> test.lin(R,co, school.lm)
$Prediction
[,1]
R1 0.6466691
R2 -0.4384634
R3 -0.4450893
$`Ho Value`
[,1]
[1,] 0
[2,] 0
[3,] 0
$`Test Stat`
[,1]
[1,] 341.2251
$`P-value`
[,1]
[1,] 1.276346e-111
There is a "canned" way to perform this hypothesis test (but not all linear hypothesis tests, read on). Namely, you could estimate the reduced model (i.e., the one where beta1=beta2=beta3=0), and then use the
anova() command to compare the two models via a likelihood ratio test. This gives an identical test statistic.
> ## Alternative method: Estimate reduced model
> ## and use anova()
>
> school.red = lm(math_scr~enrl_tot,data=caschool.df)
>
> anova(school.lm, school.red)
Analysis of Variance Table
Model 1: math_scr ~ avginc + str + meal_pct + enrl_tot
Model 2: math_scr ~ enrl_tot
Res.Df RSS Df Sum of Sq F Pr(>F)
1 415 41988
2 418 145559 3 103571 341.23
But, the value in my function is that it works for linear restrictions for which you cannot specify a full-model vs. reduced-model relationship. For example, you may want to test a more complicated null hypothesis:
Ho: beta1+beta2+beta3 =1 and beta4-beta3 = 0This is simple to specify in my framework, but it is a little hard to figure out what reduced model will give you these joint restrictions. Therefore, using
anova() isn't likely to get you very far. Here's how to implement this test using my code (code with output).
> R1 = c(0,1,1,1,0)
> R2 = c(0,0,0,-1,1)
>
> R = rbind(R1,R2)
>
> co = matrix(c(1,0), nrow=2)
>
> test.lin(R,co,school.lm)
$Prediction
[,1]
R1 -0.2368836
R2 0.4449859
$`Ho Value`
[,1]
[1,] 1
[2,] 0
$`Test Stat`
[,1]
[1,] 156.0517
$`P-value`
[,1]
[1,] 2.909143e-51
As I mentioned at the outset, I think there must be functions that do what my function does. I just didn't conduct a search for such a function (In fact, I might have used one such function in the past). Feel free to share if you know the name and package of such a function.
UPDATE: I should have mentioned this in this post before now, but I discovered a much more user-friendly method for conducting linear hypothesis tests in the car package: linearHypothesis() in car.
Here is a video tutorial where I show how to use linearHypothesis.