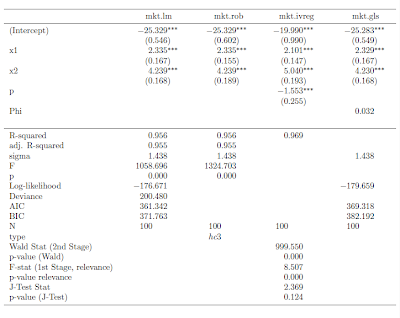
I recently discovered the mtable() command in the memisc library and its use with toLatex() to produce nice summary output for lm and glm objects in a nicely formatted table like this:
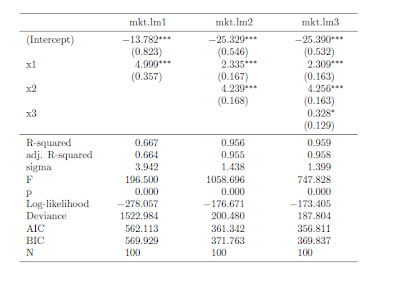
Once you have your linear model objects, all you need is one command -- a composite of toLatex() and mtable(). Here's the R code I used to generate the LaTeX code:
All you need to do from here is to copy the code into your LaTeX document (or in my case LyX document with
evil red text), where you use the dcolumn and booktabs packages (specify this in the preamble using \uspackage{} for each package).
This is really straightforward ... as long as you just want to use lm or glm objects with no deviations from that baseline. But, what if you -- as many econometricians do -- want to estimate your linear model with heteroskedasticity-consistent standard errors?
How to do it with "robust" standard errorsMy favorite way to robustify my regression in R is to use some code that John Fox wrote (and I found in an R-help forum). He called it summaryHCCM.lm(). In my own applications, I have renamed it summaryR() because "R" makes me think "robust" and it is fewer keystrokes than HCCM. I modified the output of the code to work better with what follows:
Suppose I would like to produce nicely-typed output like the above table, but with robust standard errors. Unfortunately, the summaryR() solution doesn't map naturally into our mtable() solution to producing nice LaTeX output. But, fortunately, there is a solution.
(If you just want this to work, copy and run all of the code from here to "so we are all set." No need to understand the implementation details. If you're interested in understanding how I arrived at my solution, here come the details)
It turns out that there is a way to map the summaryR() command to the mtable() command to produce nice LaTeX output with robust standard errors. Once we have the commands, the syntax is just as easy as using summaryR() directly
and it makes creating LaTeX table output easy. So, what do we do?
First, we need to adapt the methods behind mtable() to allow it to recognize something that I call a "robust" object. To do this, we need to write a function called getSummary.robust()
and we need to set some formatting options using the setSummaryTemplate() command:
Second, we need to convert a linear models object into something called a "robust" object. The robust() command does this:
So, we are all set. If you run all of the preliminary code in this post, you're in good shape to produce LaTeX tables with robust standard errors like this table:
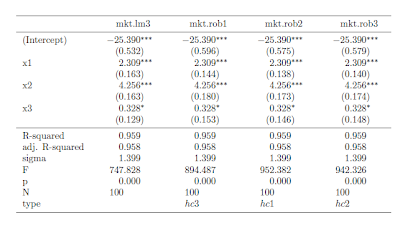
Once the functions are defined, the syntax is straightforward. Here is the code I used to put together the the table (plus, one step to copy the output into my LyX file)
I hope this is helpful. If you're interested in other extensions of mtable(),
here's an excellent post.
Finally,
I am still learning I learned how to bundle functions into a package (so I have
n't made this available as a package
yet). Today, I was able to successfully package this set of functions into a package called tonymisc on my computer.
Once I organize the help/example/other fun parts of the package, I will see about making this and my extension for ivreg objects available. This package (in its first version) is available on CRAN through install.packages("tonymisc").